Exploring fungal orthologs with a human nucleolar proteins
This page expands the yeast complex-based utility demonstration by using a broader human nucleolar protein set (n = 94) curated from The Human Protein Atlas. Rather than focusing on a complex, this utility page asks a complementary question: how widely are human nucleolar proteins represented as fungal orthologs in FuNGI, and how often do those orthologs show predicted NoLS/NLS support?
It first provides a global proteome-level overview, then allows in-depth exploration of one selected fungal proteome and the fungal orthologs corresponding to 94 human nucleolar proteins.
FuNGI catalogs only proteins with at least one predicted NoLS. Therefore, even if an ortholog detected in a fungal proteome, it may not be displayed in the database unless it contains predicted NoLSs.
Proteome-level overview
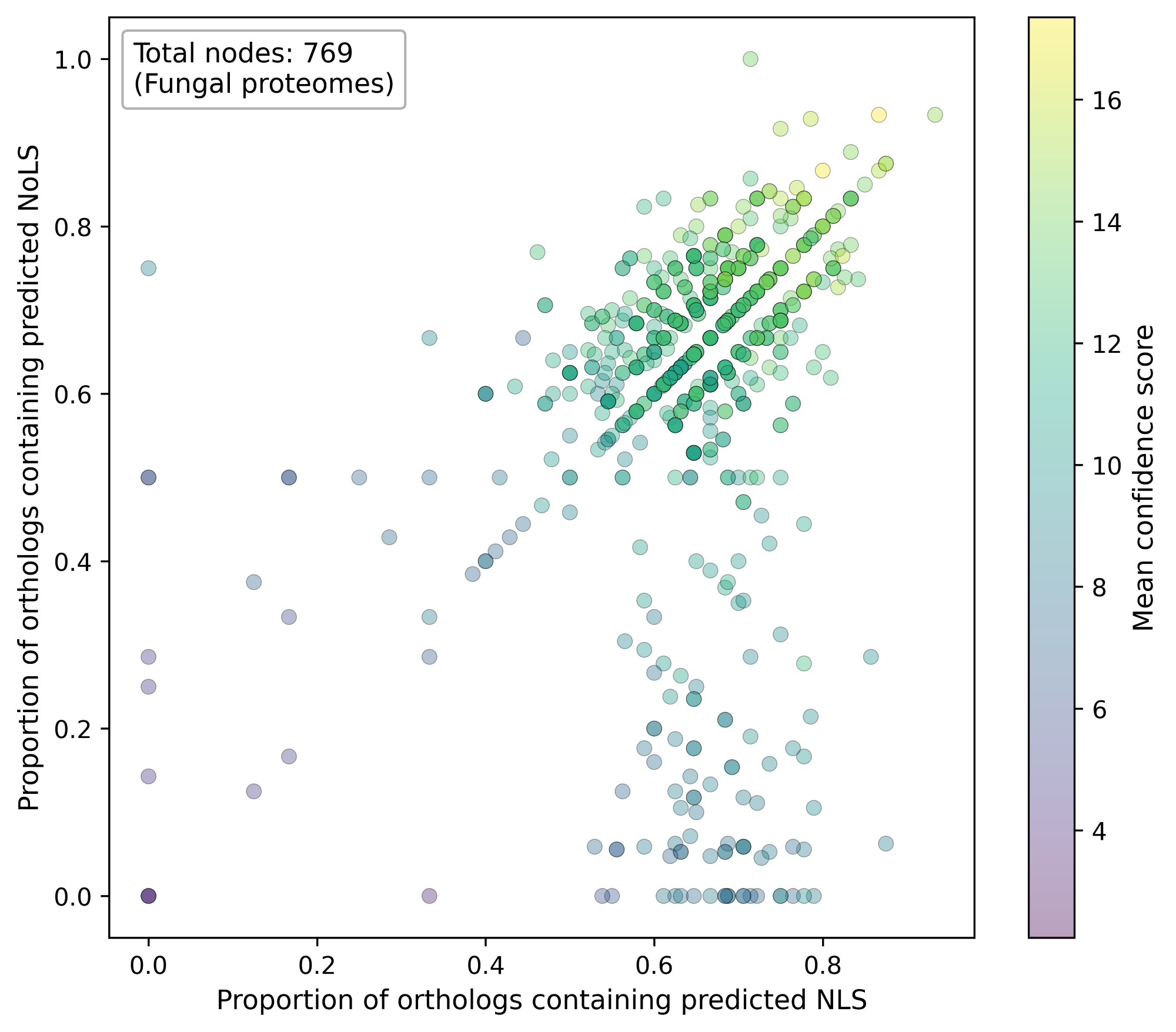
Each point represents one fungal proteome. The x-axis is the proportion of detected orthologs containing predicted NLS, and the y-axis is the proportion of detected orthologs containing predicted NoLS.
Point color represents the mean FuNGI confidence score of detected orthologs.
At the proteome level, substantial proportions of fungal orthologs derived from the 94 human nucleolar proteins contain predicted NoLSs and NLSs and are consistently assigned medium-to-high confidence scores in FuNGI.
Fungal orthologs of the selected proteome
| Query (Human Protein) | Subject (Fungal Ortholog) | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Reference protein | UniProt | Ortholog | Gene name | # NoLS | # NLS | PSORT | Confidence | RBH | UniProt |
| sp|O15213|WDR46_HUMAN | O15213 | Q6CTX7 | KLLA0_C09262g | 2 | 2 | nucl | High (15.9) |
F: 43.154 / 75 R: 42.887 / 85 |

|
| sp|O15381|NVL_HUMAN | O15381 | Q6CQ05 | KLLA0_E00837g | 1 | 1 | nucl | Medium (13.8) |
F: 51.475 / 69 R: 51.475 / 74 |

|
| sp|Q13206|DDX10_HUMAN | Q13206 | Q6CRF4 | DBP4 | 3 | 1 | nucl | High (19.5) |
F: 49.704 / 57 R: 53.691 / 58 |

|
| sp|Q15050|RRS1_HUMAN | Q15050 | Q6CVC7 | KLLA0_B13046g | 1 | 1 | nucl | Medium (13.4) |
F: 38.22 / 47 R: 38.22 / 93 |

|
| sp|Q15397|PUM3_HUMAN | Q15397 | Q6CTJ1 | KLLA0_C12331g | 2 | 0 | nucl | Medium (11.7) |
F: 27.5 / 82 R: 27.355 / 81 |

|
| sp|Q5C9Z4|NOM1_HUMAN | Q5C9Z4 | Q6CW80 | KLLA0_B06116g | 2 | 1 | nucl | High (15.5) |
F: 32.483 / 47 R: 32.483 / 40 |

|
| sp|Q96GQ7|DDX27_HUMAN | Q96GQ7 | Q6CJV1 | DRS1 | 2 | 3 | nucl | High (19.5) |
F: 44.118 / 61 R: 45.77 / 61 |

|
| sp|Q9BRU9|UTP23_HUMAN | Q9BRU9 | Q6CSH3 | KLLA0_D01023g | 2 | 2 | nucl | High (18.5) |
F: 29.954 / 85 R: 30.263 / 84 |

|
| sp|Q9BSC4|NOL10_HUMAN | Q9BSC4 | Q6CXY5 | KLLA0_A04653g | 3 | 2 | nucl | High (20.5) |
F: 44.902 / 79 R: 40.56 / 94 |

|
| sp|Q9H0S4|DDX47_HUMAN | Q9H0S4 | Q6CT85 | RRP3 | 1 | 1 | nucl | Medium (14.4) |
F: 54.81 / 95 R: 54.81 / 92 |

|
| sp|Q9H9Y2|RPF1_HUMAN | Q9H9Y2 | Q6CJ42 | KLLA0_F21670g | 1 | 1 | nucl | Medium (14.4) |
F: 47.811 / 84 R: 46.801 / 98 |

|
| sp|Q9NWT1|PK1IP_HUMAN | Q9NWT1 | Q6CP30 | KLLA0_E07965g | 1 | 1 | cyto_nucl | Medium (10.8) |
F: 27.761 / 76 R: 27.513 / 91 |

|
What this utility shows
This utility demonstrates that FuNGI can be explored not only through well-characterized complexes, but also through a broader cross-species reference proteins.
By starting from 94 human nucleolar proteins and identifying their fungal orthologs, users can explore, within a fungal proteome of interest, how strongly individual orthologs exhibit predicted nucleolar localization signals and are prioritized by FuNGI’s integrated prediction framework.
Together with the yeast complex-based demo, this page shows that FuNGI captures a substantial fraction of conserved nucleolar protein orthologs across fungal proteomes, suggesting its utility as a prioritization framework for identifying strongly supported nucleolar localization signals.