Welcome to FuNGI!
Fungal Nucleolar Genomic Inventory —
A Home for Fungal Proteins with Predicted Nucleolar Localization Signals
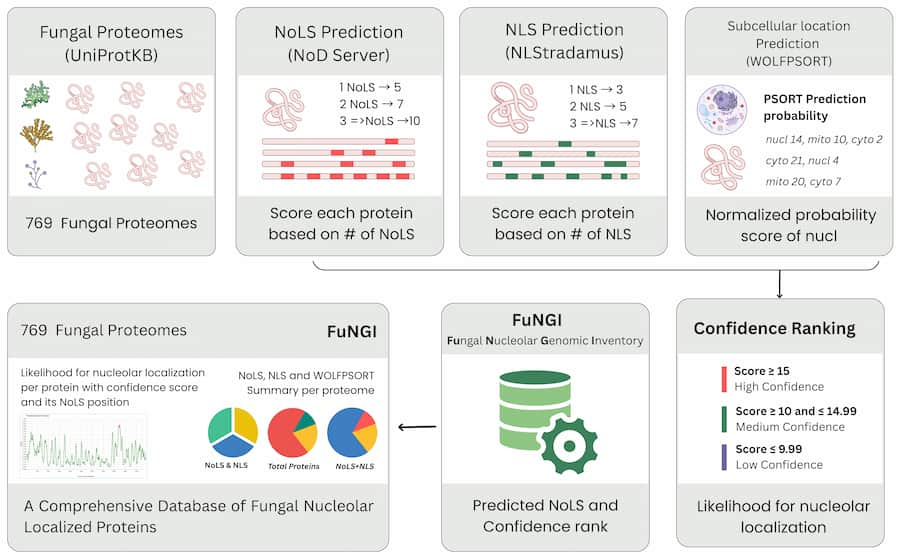
Fungal Nucleolar Genomic Inventory —
A Home for Fungal Proteins with Predicted Nucleolar Localization Signals

| # Proteomes | 769 |
|---|---|
| # NoLS+ | 1,729,729 |
| # NLS+ | 1,437,299 |
| # NoLS+&NLS+ | 985,501 |
| Avg. NoLS+ | 2,249 (per proteome) |
| Top Localization | Cyto, Nucleus, Mito, Cyto_Nucleus |
Fungal proteins with predicted nucleolar localization signals across hundreds of fungal proteomes, enabling large-scale comparative analysis.
Multiple ways to access and analyze nucleolar localization signal data across fungal species through taxonomy and proteome-specific views.
Proteins are ranked using a scoring scheme that reflects their likelihood of nucleolar localization, facilitating prioritization for future experimental validation.
Experimentally validated nucleolar localization proteins from M. oryzae, demonstrating the predictive reliability of FuNGI.